AFM Systems
AFM Accessories
Learning
Contact Us
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
A mitochondrion – the powerhouse of a cell – harbors its own DNA genome. Mutations and deletions in this genome are correlated with numerous neuromuscular diseases and premature aging. Thus, a detailed understanding of how mitochondrial DNA undergoes replication, repair, RNA translation, and more could provide important insights into these diseases and their treatments. This is currently one area of research focus for Parminder Kaur, a Research Professor in Physics at North Carolina State University (NCSU).
The replication of mitochondrial DNA starts in the same manner as all double-stranded DNA. A large protein complex called a helicase separates (unwinds) the two strands analogously to the way a zipper is pulled open.
In mammalian mitochondria, the particular type of helicase is called “Twinkle.” One of the interesting novelties of Twinkle, besides its fanciful name, is the mystery of how it engages with DNA without the presence of a “loader” protein. These loader proteins are known to be a necessary co-factor in DNA replication in the cell nucleus.
A collaborative study between Professor Kaur and Professor Hong Wang at NCSU, together with researchers at Rice University has recently made substantial progress in unravelling some details of the mechanism [i]. Professor Kaur acquired some of the key dynamic data in the form of AFM videos and images.

|
“The Cypher VRS combines three vitally important advantages for this research: high speed, stability under liquid, and a large field of view.” — Professor Parminder Kaur, Department of Physics, North Carolina State University. |
Professor Kaur explains, “Twinkle is a lock-washer-shaped hexamer; it consists of six distinct domains arranged in a ring format with very flexible links between the domains. In simple terms we wanted to use AFM imaging in real-time to investigate how this closed ring binds with a single-stranded region of DNA and then slides along the double-strand.”
Her collaborators at Rice had used cryogenic electron microscopy (cryo-EM) to get static images of Twinkle. However, like many other multi-domain protein complexes, a significant amount of Twinkle is disordered and dynamic. This can’t be accurately imaged in cryo-EM because of the averaging process involved.
Instead, Professor Kaur turned to the unique imaging capabilities of an AFM with video-rate speed (VRS) – specifically the Cypher VRS. And while other researchers have previously obtained AFM data of protein complexes interacting with membrane-bound DNA, no-one had successfully imaged a hexamer interacting with free DNA in a liquid environment.
Using the Cypher VRS, her group obtained high-resolution videos at 1-2 frames/second. The results were quite surprising. When Twinkle is in the proximity of single stranded DNA, it undergoes a dramatic conformational change. The so-called N domain reaches out and appears to capture the DNA in an arm-like motion (see figure). They also looked at the impact of Twinkle mutations on this behavior.
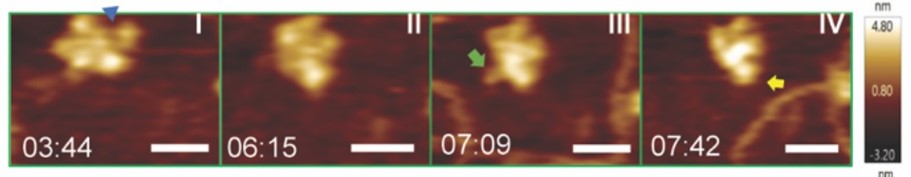
Real-time HS-AFM imaging of LcTwinkle shows DNA capture by N-protrusion and at the central channel. A. LcTwinkle switches between open (blue triangle) and closed conformations in the absence of DNA (Panels I and II). Nearby DNA induces LcTwinkle N-protrusion (Panels III and IV).
Although Professor Kaur and her colleagues believe Twinkle must be responding to some type of electrostatic force, they were surprised that Twinkle clearly is able to sense DNA even at distances of over 10 nm and tries to reach towards the DNA. Together with the cryo-EM data, these videos enabled the team to draw several important conclusions about the way DNA replication is performed and regulated in mitochondria.
Professor Kaur explains the enabling role of the Cypher VRS. She states, “It combines three vitally important advantages for this research: high-speed, stability under liquid, and a large field of view. We needed to record videos at 1-2 frames per second to see the capture process happening.”
She notes that the Cypher VRS, in particular, avoids the traditional AFM tradeoff between speed and field of view. This enabled her to image at high frame rates with fields as large as 1 µm x 1 µm. She could then track multiple Twinkle molecules at the same time, which was necessary to randomly observe the capture process happening with at least one of them.
Professor Kaur also points out the importance of the probe stability and benign tapping resulting from photothermal (blueDrive) oscillation. “I have a lot of experience in using AFMs to look at single biomolecules under water. A limitation of mechanical (piezo) cantilevers in this situation is that they do not produce a clean oscillation under liquids. This means the sample and DNA motion could couple into vibration of the cantilever. We had no such limitations with Cypher. We could clearly see the DNA in motion even at 1 frame/sec, without it getting lost in any vibration of the cantilever. I highly recommend photothermal oscillation for imaging at high speed under buffer environments to capture biomolecules and specifically DNA in action.”
[i] Li, Z., Kaur, P., Lo, C.Y., Chopra, N., Smith, J., Wang, H. and Gao, Y., 2022. Structural and dynamic basis of DNA capture and translocation by mitochondrial Twinkle helicase. Nucleic Acids Research, 50(20), pp.11965-11978.
Author: Asylum Research
Category: Case Study
