AFM Systems
AFM Accessories
Learning
Contact Us
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Researchers in China and the U.S. studied polymerization of curli fibrils using DNA origami as molecular nucleators. High-resolution, video-rate AFM topography imaging revealed details of fibril formation dynamics on the single-molecule level.


Amyloid fibrils are protein aggregates that perform a wide range of biological functions. An example are curli fibrils, functional amyloids present in biofilms of E. coli bacteria. It is known that cells often use molecular-scale nucleators to mediate formation of these fibrils to ensure proper structure and function. However, many details of this process are not clearly understood.
In this work, a team led by researchers at ShanghaiTech University gained deeper insight into fibril formation using AFM techniques. They decorated DNA origami with curli-specific gene B proteins to mimic protein nucleators. High-speed topography movies allowed direct visualization of fibril assembly dynamics on the single-molecule level, while closeup topography images gave further insight. The experiments revealed that fibril growth was stimulated by the curli-decorated origami in two distinct ways.
The results provide an in-vitro platform for probing how nucleators regulate fibril polymerization that could benefit a range of applications in bionanotechnology and medicine.
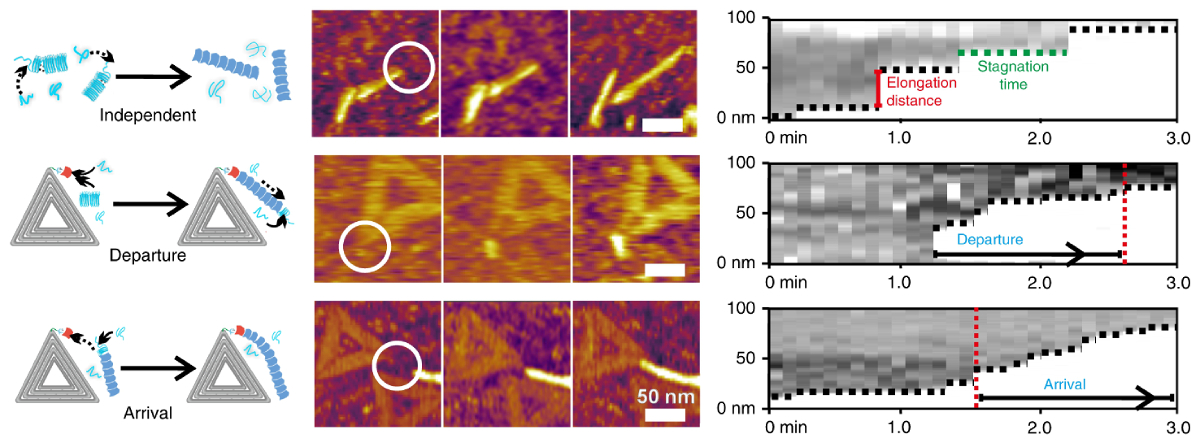
Movies of nanoscale topography were acquired with tapping mode in liquid on the Cypher VRS AFM. Time-lapse images from the movies are shown in the main paper, while the entire movies are available in the supplementary information at the journal website. At about 6 seconds per frame, the movies illustrate the power of the VRS for capturing nanoscale dynamic processes with the resolution and ease of standard Cypher AFMs. Additional topography images were acquired on the MFP-3D AFM with tapping mode in liquid and air to more closely examine fibril and origami structure.
Citation: X. Mao, K. Li, M. Liu et al. Directing curli polymerization with DNA origami nucleators. Nat. Commun. 10, 1395 (2019). https://doi.org/10.1038/s41467-019-09369-6
Note: The original article featured above was published as Open Access. The data shown here are reused under fair use of the original article and under a Creative Commons license, which can be accessed through the article link above.
Author: Asylum Research
Category: Technical Article
